APOGEE Spectra with Bayesian NN and Gaia offset calibration - ApogeeDR14GaiaDR2BCNN
- class astroNN.models.apogee_models.ApogeeDR14GaiaDR2BCNN(lr=0.001, dropout_rate=0.3)[source]
Class for Bayesian convolutional neural network for APOGEE DR14 Gaia DR2
- History
2018-Nov-06 - Written - Henry Leung (University of Toronto)

ApogeeDR14GaiaDR2BCNN can only be used with Apogee spectra with 7,514 pixels
from astroNN.models import ApogeeDR14GaiaDR2BCNN
from astroNN.datasets import H5Loader
# Load the train data from dataset first, x_train is spectra and y_train will be ASPCAP labels
loader = H5Loader('datasets.h5')
loader.load_combined = True
loader.load_err = False
loader.target = ['Ks-band fakemag']
x_train, y_train, x_err, y_err = loader.load()
# And then create an instance of Apogee Censored Bayesian Convolutional Neural Network class
apogee_gaia_bcnn = ApogeeDR14GaiaDR2BCNN()
# Set max_epochs to 10 for a quick result. You should train more epochs normally, especially with dropout
apogee_gaia_bcnn.max_epochs = 10
apogee_gaia_bcnn.train(x_train, y_train, x_err, y_err)
Here is a list of parameter you can set but you can also not set them to use default
ApogeeDR14GaiaDR2BCNN.batch_size = 64
ApogeeDR14GaiaDR2BCNN.initializer = 'he_normal'
ApogeeDR14GaiaDR2BCNN.activation = 'relu'
ApogeeDR14GaiaDR2BCNN.num_filters = [2, 4]
ApogeeDR14GaiaDR2BCNN.filter_len = 8
ApogeeDR14GaiaDR2BCNN.pool_length = 4
# number of neurone for [old_bcnn_1, old_bcnn_2, offset_hidden_1, offset_hidden_2]
ApogeeDR14GaiaDR2BCNN.num_hidden = [162, 64, 32, 16]
ApogeeDR14GaiaDR2BCNN.max_epochs = 50
ApogeeDR14GaiaDR2BCNN.lr = 0.005
ApogeeDR14GaiaDR2BCNN.reduce_lr_epsilon = 0.00005
ApogeeDR14GaiaDR2BCNN.reduce_lr_min = 0.0000000001
ApogeeDR14GaiaDR2BCNN.reduce_lr_patience = 10
ApogeeDR14GaiaDR2BCNN.target = 'all'
ApogeeDR14GaiaDR2BCNN.l2 = 5e-9
ApogeeDR14GaiaDR2BCNN.dropout_rate = 0.2
ApogeeDR14GaiaDR2BCNN.input_norm_mode = 3
ApogeeDR14GaiaDR2BCNN.labels_norm_mode = 2
Note
You can disable astroNN data normalization via ApogeeDR14GaiaDR2BCNN.input_norm_mode=0 as well as ApogeeDR14GaiaDR2BCNN.labels_norm_mode=0 and do normalization yourself. But make sure you don’t normalize labels with MAGIC_NUMBER (missing labels).
After the training, you can use apogee_gaia_bcnn in this case and call test method to test the neural network on test data. Or you can load the folder by
from astroNN.models import load_folder
apogee_gaia_bcnn = load_folder('astroNN_0101_run001')
# Load the test data from dataset, x_test is spectra and y_test will be ASPCAP labels
test_data = ......
# pred contains denormalized result aka. fakemag prediction in this case
# pred_std is a list of uncertainty
# pred_std['total'] is the total uncertainty (standard derivation) which is the sum of all the uncertainty
# pred_std['predictive'] is the predictive uncertainty predicted by bayesian neural net
# pred_std['model'] is the model uncertainty from dropout variational inference
pred, pred_std = apogee_gaia_bcnn.test(test_data)
# Calculate jacobian
jacobian_array = apogee_gaia_bcnn.jacobian(x_test, mean_output=True)
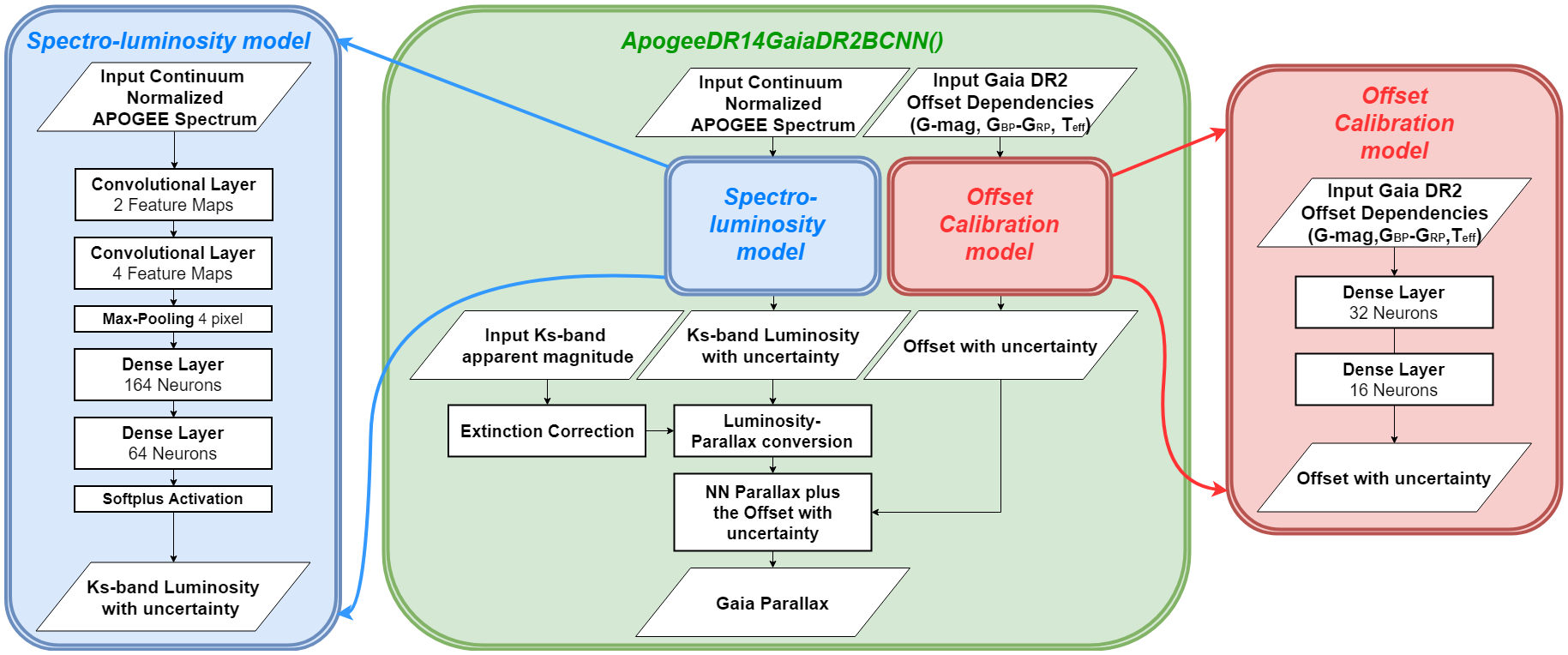
Architecture
The architecture of this neural network is as follow.