Compiling and Loading APOGEE and Gaia Datasets - astroNN.datasets¶
Compiling APOGEE Dataset¶
from astroNN.datasets import H5Compiler
# To create a astroNN compiler instance
compiler = H5Compiler()
# To set the name of the resulting h5 datasets, here a 'test.h5' will be created
compiler.filename = 'test'
# To compile a .h5 datasets, use .compile() method
compiler.compile()
# Avaliable attributes of astroNN H5Compiler, set them before using H5Compiler.compiler()
H5Compiler.apogee_dr # APOGEE DR to use, Default is 14
H5Compiler.gaia_dr # Gaia DR to use, Default is 1
H5Compiler.starflagcut = True # True to filter out ASPCAP star flagged spectra
H5Compiler.aspcapflagcut = True # True to filter out ASPCAP flagged spectra
H5Compiler.vscattercut = 1 # Upper bound of velocity scattering
H5Compiler.teff_high = 5500 # Upper bound of SNR
H5Compiler.teff_low = 4000 # Lower bound of SNR
H5Compiler.SNR_low = 200 # Lower bound of SNR
H5Compiler.SNR_high = 99999 # Upper bound of SNR
H5Compiler.ironlow = -3 # Lower bound of SNR
H5Compiler.filename = None # Filename of the resulting .h5 file
H5Compiler.spectra_only = False # True to include spectra only without any aspcap abundances
H5Compiler.cont_mask = None # Continuum Mask, none to use default mask
H5Compiler.use_apogee = True # Currently no effect
H5Compiler.use_esa_gaia = True # True to use ESA Gaia parallax, **if use_esa_gaia is True, ESA Gaia will has priority over Anderson 2017**
H5Compiler.use_anderson_2017 = False # True to use Anderson et al 2017 parallax, **if use_esa_gaia is True, ESA Gaia will has priority**
H5Compiler.err_info = True # Whether to include error information in h5 dataset
H5Compiler.continuum = True # True to do continuum normalization, False to use aspcap normalized spectra
As a result, test.h5 will be created as shown below. you can use H5View to inspect the data
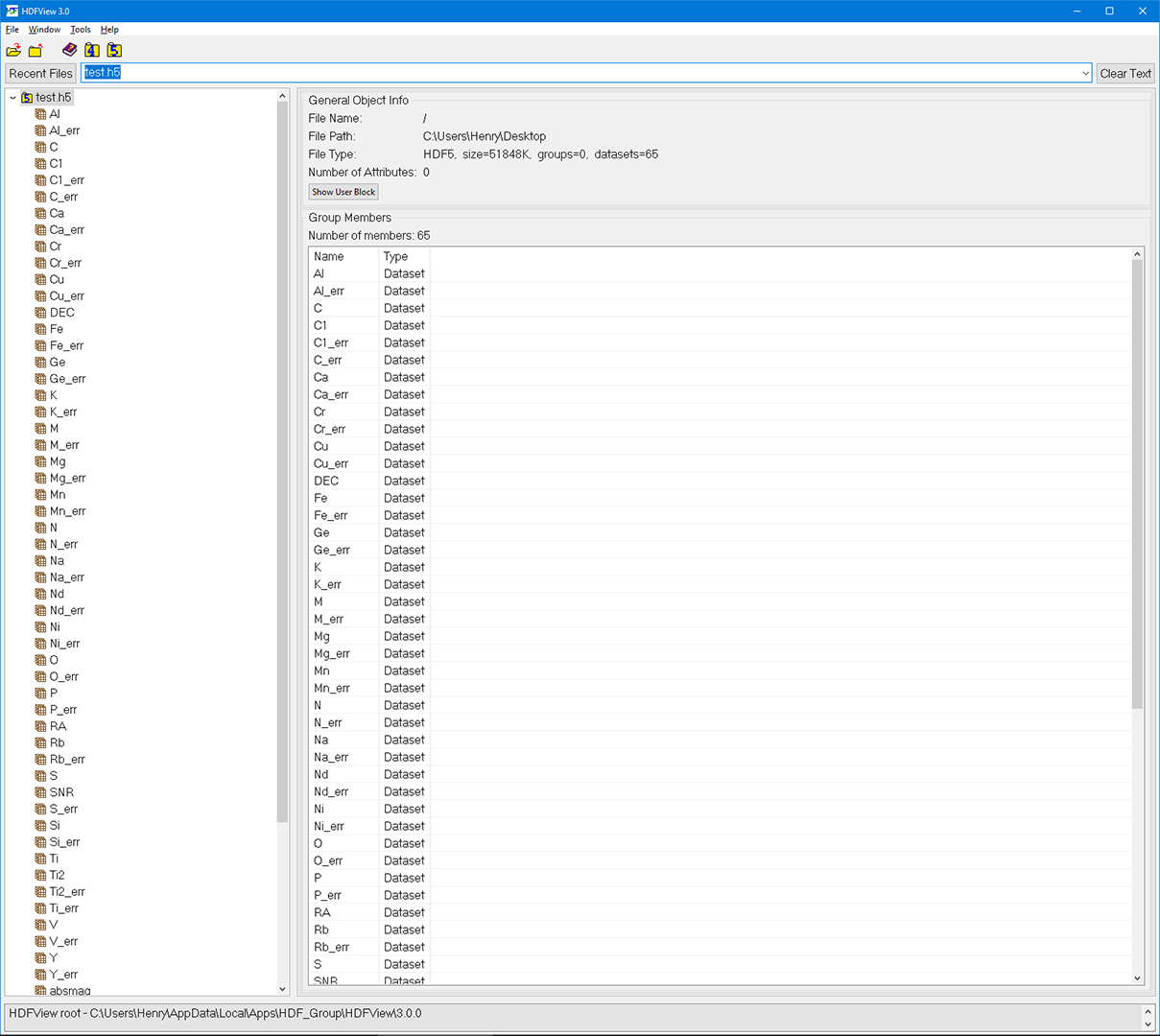
Note
For more detail on L. Anderson et al. (2017) improved parallax using data-driven stars model: arXiv:1706.05055
Loading APOGEE Dataset¶
To load a compiled dataset, you can use
from astroNN.datasets import H5Loader
loader = H5Loader('datasets.h5') # You should replace datasets.h5 with your real filename
x, y = loader.load()
loader.load_err = True # load error info too
x, y, x_err, y_err = loader.load()
# Lets say you want to load the corresponding SNR, apparent magnitude and coordinates of the spectra loaded previously
snr = loader.load_entry('SNR')
kmag = loader.load_entry('Kmag')
ra = loader.load_entry('RA')
dec = loader.load_entry('DEC')
x will be an array of spectra [training data] and y will be an array of ASPCAP labels [training labels]
#Avaliable attributes of astroNN H5Loader, set them before H5Loader.load()
H5Loader.load_combined = True # Whether to load combined spectra or individual visits
#Target 'all' means ['teff', 'logg', 'M', 'alpha', 'C', 'C1', 'N', 'O', 'Na', 'Mg', 'Al', 'Si', 'P', 'S', 'K', 'Ca', 'Ti', 'Ti2', 'V', 'Cr',
#'Mn', 'Fe', 'Co', 'Ni', 'fakemag']
H5Loader.target = 'all'
# Whether to exclude all spectra contains -9999 in any ASPCAP abundances, By default, astroNN can handle -9999 in training data
H5Loader.exclude9999 = False
# Whether to load error data
H5Loader.load_err = True
# True to load combined spectra, False to load individual visit (If there is any in the h5 dataset you compiled)
# Training on combined spectra and test on individual spectra is recommended
H5Loader.load_combined = True
You can also use scikit-learn train_test_split to split x and y into training set and testing set.
In case of APOGEE spectra, x_train and x_test are training and testing spectra. y_train and y_test are training and testing ASPCAP labels
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.2)